Discover the department
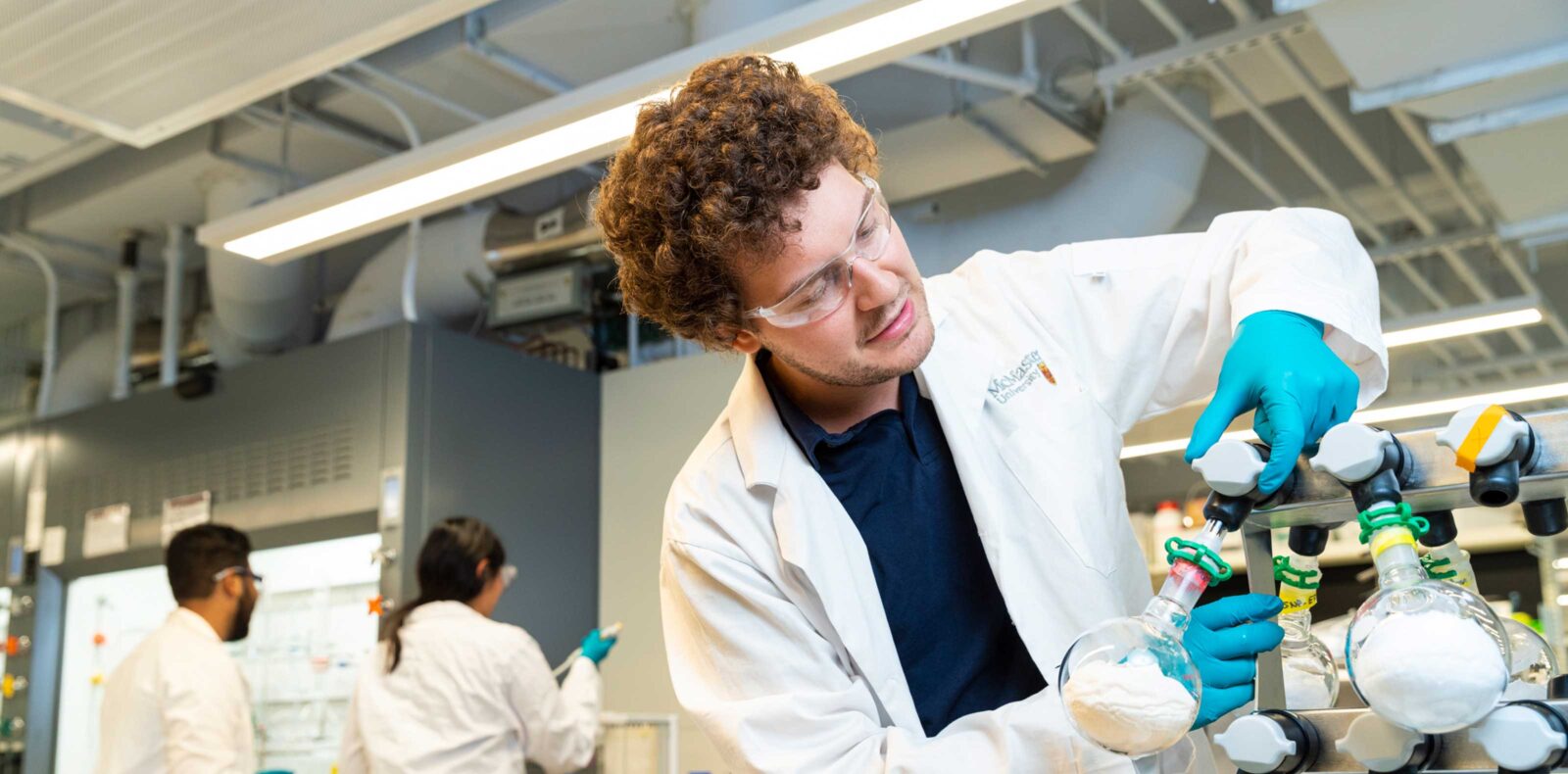
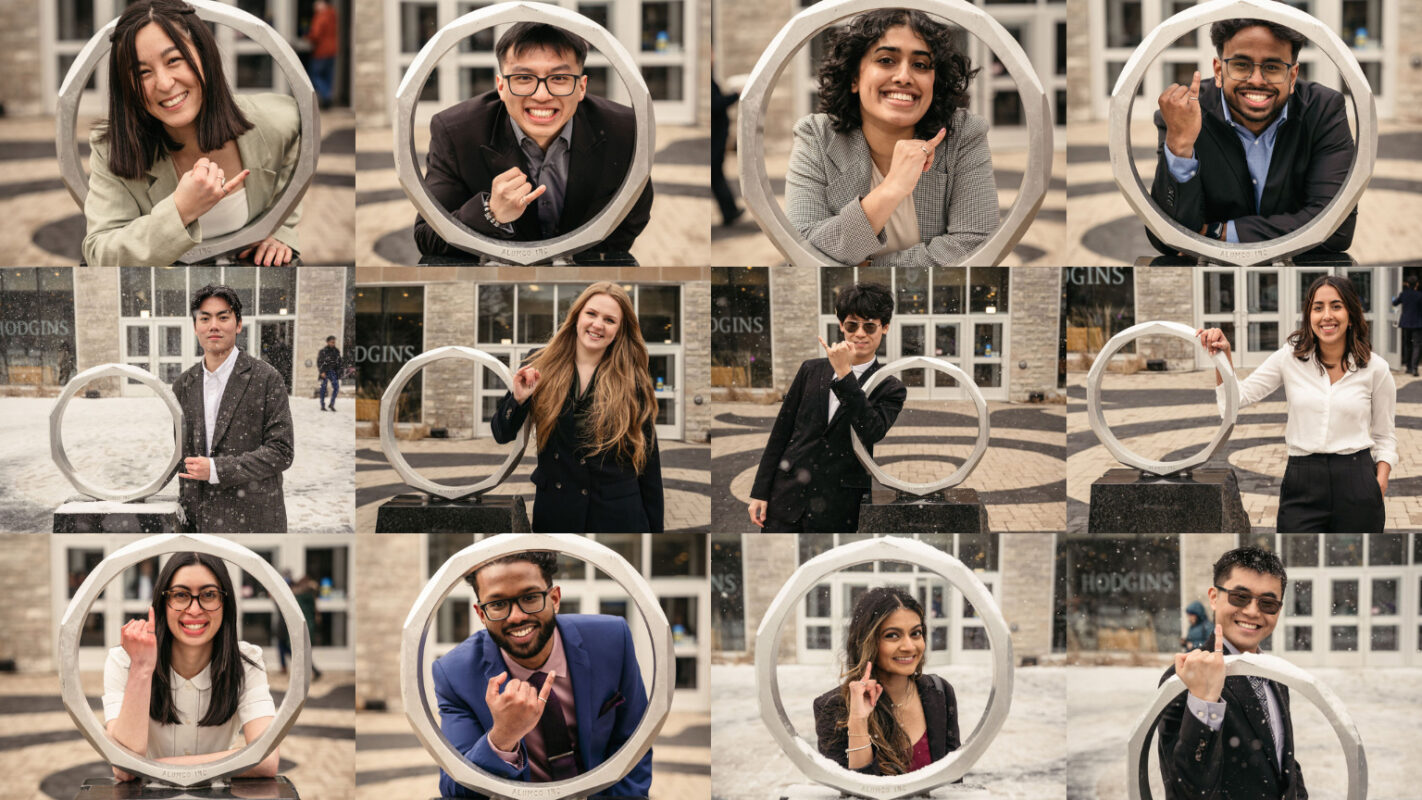
Our award-winning excellence in teaching and research will give you an edge whether you’re starting or expanding your career. Our dynamic program will give you the very best training in this exciting and ever-evolving discipline.
-
57%
women enrolled
-
4
Canada Research Chairs
-
83%
of graduating undergraduates had a co-op experience
What’s happening
There are no upcoming events scheduled for this area. Please check back soon for more information.